samsam documentation#
The samsam package provides two samplers:
samsam.sam : a Scaled Adaptive Metropolis algorithm (see [1], [2], [3]), to robustly obtain samples from a target distribution,
samsam.covis : a COVariance Importance Sampling algorithm, to efficiently compute the model evidence (or other integrals).
It additionally includes tools (samsam.acf) to assess the convergence of the sam sampler (ACF, IAT), and a few commonly used prior distributions (samsam.logprior).
Installation#
Using conda#
The samsam package can be installed using conda with the following command:
conda install -c conda-forge samsam
Using pip#
It can also be installed using pip with:
pip install samsam
Example#
Let us first define a simple log-probability function:
In [1]: import numpy as np
...: import matplotlib.pyplot as plt
...: from samsam import sam, covis, acf
...: from corner import corner
...: np.random.seed(0)
...:
...: def logprob(x):
...: return(-0.5*(np.sum(x**2) + x.size*np.log(2*np.pi)))
...:
Then we run sam to sample this distribution:
In [2]: ndim = 10
...: nsamples = 100000
...: x0 = np.random.normal(0, 100, ndim)
...:
...: samples, sam_diagnos = sam(x0, logprob, nsamples=nsamples, print_level=0)
...: samples = samples[nsamples//4:]
...:
Let us check that sam converged correctly using the ACF/IAT:
In [3]: R = acf.acf(samples)
...: tau = np.arange(samples.shape[0])
...:
...: plt.figure()
...: plt.plot(tau[1:], R[1:])
...: plt.xscale('log')
...: plt.xlim(1, samples.shape[0])
...: plt.xlabel('lag')
...: plt.ylabel('ACF')
...:
...: iat = acf.iat(R=R)
...: print('IAT:', iat.max())
...: print('Effective number of samples:', samples.shape[0]/iat.max())
...:
IAT: 40.59274361034297
Effective number of samples: 1847.6454984158759
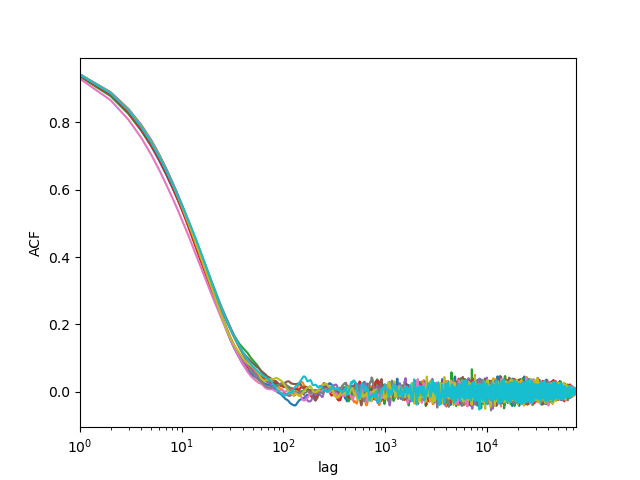
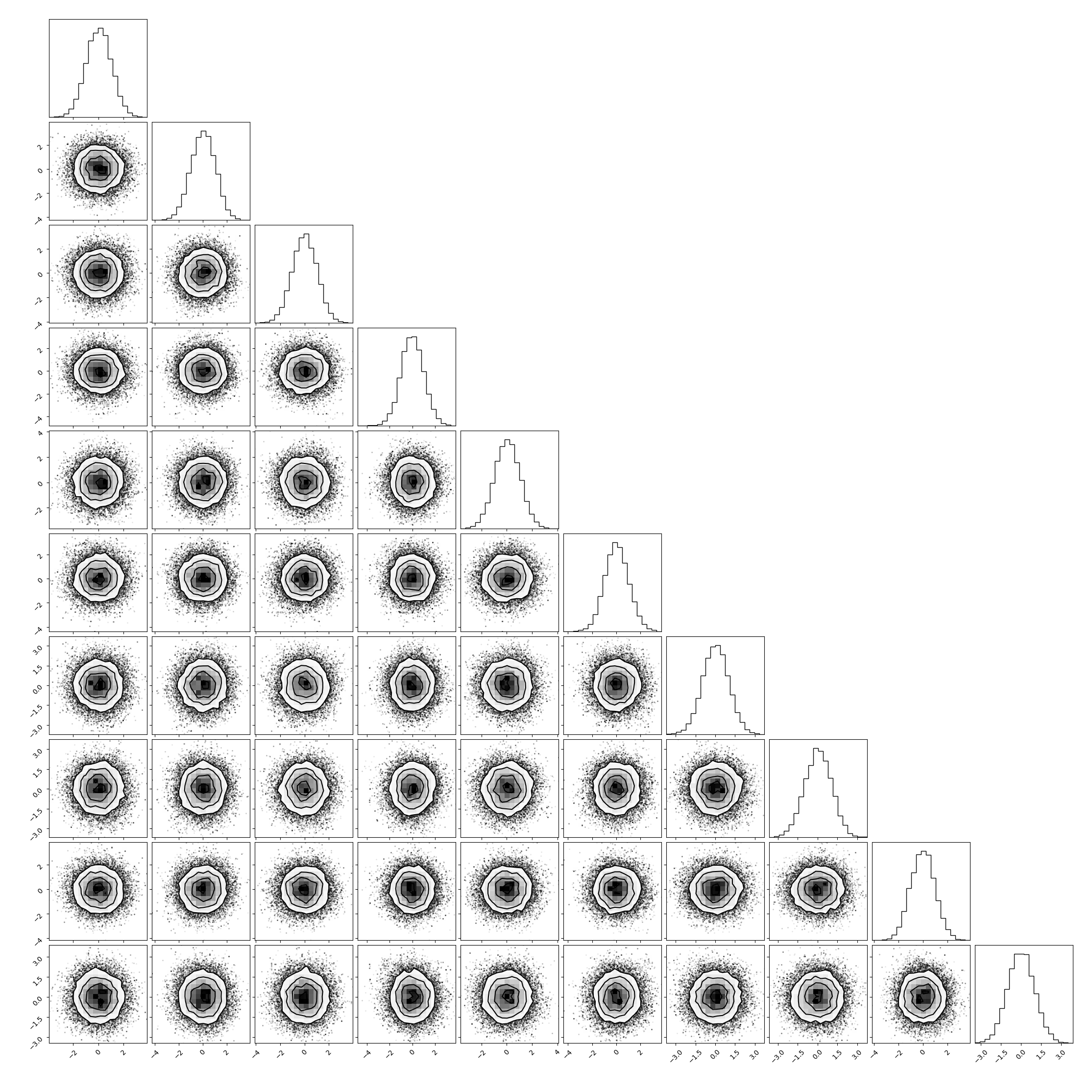
Now we plot the corner plot of the parameter samples:
In [4]: corner(samples);

Finally we run covis to compute the log-evidence of the model:
In [5]: _, _, covis_diagnos = covis(sam_diagnos['mu'], sam_diagnos['cov'], logprob, nsamples=1000, print_level=0)
...: # Should be close to 0 since the logprob is correctly normalized
...: print('Log-evidence:', covis_diagnos['logevidence'])
...:
Log-evidence: -0.002628219768561202